Unlocking microbial functionality to understand the impact of functional ingredients



As consumers continue to turn towards more natural ways to maintain their health, the market for probiotics and other ingredients that promote a healthy gut microbiome continues to grow. But how can a manufacturer be sure their pre-, pro- or postbiotic will have a real impact? Advances in molecular microbiology and bioinformatics are unlocking more information from bacterial DNA and RNA, and this provides insight into the effects of ingredients on microbial functionality and human health.
The gut microbiome includes the trillions of living microorganisms in the gut, creating a complex community that impacts not only gut health, but also everything from general health to sleep, immunity, the gut-brain axis and more. The gut microbiome can be modulated by components and foods that, for example, promote ‘good’ bacteria or inhibit the ‘bad’ ones. This offers manufacturers opportunities to add value to their products, by adding pre-, pro-, and postbiotics and other ingredients that modulate the microbiome in a way that delivers a health benefit to the consumer.
If you want to add an ingredient to a product, or want to develop a product with a specific functionality, you need to understand what happens in the gut after the product has been consumed. One way to do this involves studying changes to microbiome composition and the abundance of specific (groups of) species, based on microbial metagenomics (DNA). For example, if you want to look at certain bacteria that are known to produce the short chain fatty acid (SCFA) butyrate, which offer health benefits, you can define their presence by isolating and identifying their DNA, and then determine if their relative abundance has gone up or down. However, this won’t tell you whether the bacteria are producing butyrate in this situation.
But as RNA-based techniques have become both better and more accessible, we are able to focus in on microbial functionality, instead of just composition. Functionality provides greater insight into what the bacteria are actually doing, rather than what they might do.
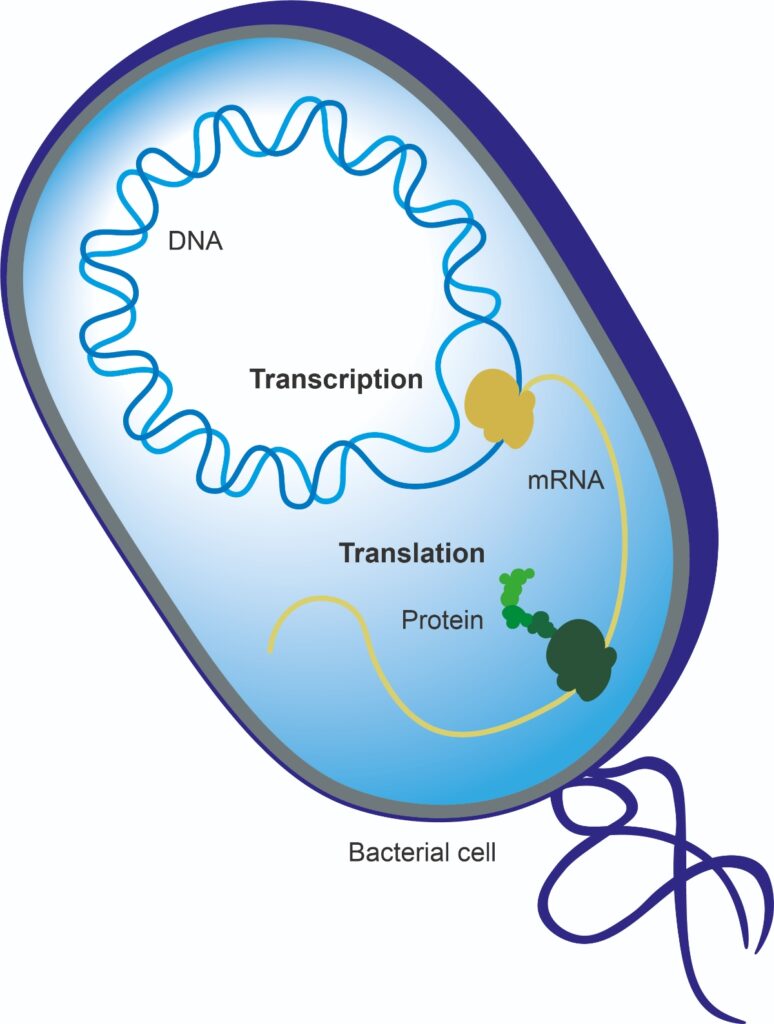
DNA and RNA are both important molecules involved in the storage and reading of the genetic information of a living being. DNA encodes, replicates and stores genetic information, and messenger RNA (mRNA) converts that information to a format that can build proteins. These proteins are the workhorses of the cell that actually deliver the functionality.
DNA can thus tell us about the potential of a bacterial species to form a protein or enzyme, while RNA provides greater insight into whether the bacteria are actually doing when, for example, a probiotic has been added to a food product and consumed.
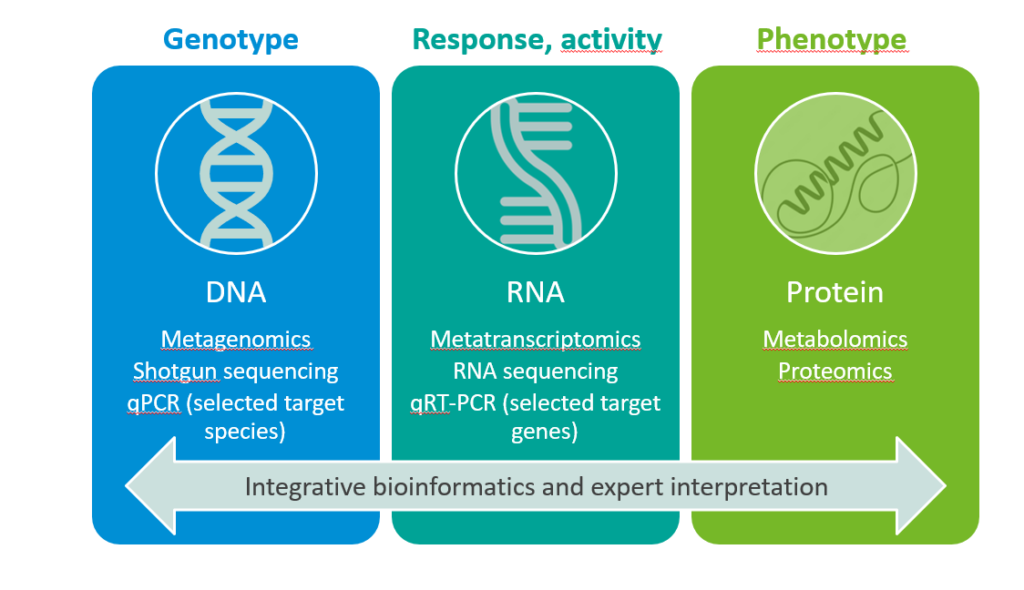
Based on samples collected from either in vitro studies or clinical trials, there are a broad range of bioinformatics methods you can run on both DNA and RNA sequencing data. Existing sources, such as generic functional annotation databases, can provide information on a gene’s biological identity. This rather limited insight can be enriched with tools such as profile Hidden Markov Models (pHMMs), which can predict specific gene functionalities of even unknown bacteria. Shotgun metagenomics sequencing is a technique that has become much more accessible in recent years; it allows the analysis of all genes of all microorganisms that are present in a sample.
RNA sequencing, on the other hand, is applied to detect and identify which genes have been copied from DNA to mRNA in a certain situation (transcriptome). By measuring gene expression and activation (i.e., whether the information encoded in the genes has actually been used to make mRNA), and comparing two or more different situations, we can better understand the microbial response to an ingredient. For specific biomarkers of interest, quantitative polymerase chain reaction (qPCR) assays can be developed and applied to both DNA and RNA, to quantitatively measure DNA ((sub)species abundance) and gene expression (functionality abundance).
Let’s take the case of a manufacturer who wants to investigate whether their food ingredient improves the digestibility of human milk oligosaccharides (HMOs). We can follow a step-by-step approach to determine whether the ingredient leads to better microbial utilisation, and thus digestibility:
First, we isolate microbial DNA from a faecal sample and perform shotgun metagenomics sequencing. This provides insight into changes in the composition of the gut microbiota, as well as in generic gene and pathway abundances. Optional in silico screening using targeted pHMM-based predictive modules lets us know whether there is an increase in certain genes encoding specific HMO-degrading enzymes, such as fucosidases and sialidases, after the ingredient has been consumed.
We then can isolate and sequence RNA to investigate which microbial genes and pathways are more or less transcribed after the ingredient has been consumed. This helps for selecting which biomarkers to use to study functionality and microbial response mechanisms. In some cases, an optional quantitative Reverse Transcription PCR (qRT-PCR) assay can be used to measure the RNA transcript of a specific gene of an interesting species or sub-species. This can offer quantitative insight into the expression of the gene associated with HMO degradation after the intervention.
Next, in vitro measurement of specific HMO metabolites can show that there indeed is active microbial HMO metabolisation, supporting the health claims with quantitative data and knowledge about the underlying molecular mechanism.
It is important to keep in mind that investigating the transcriptomic response is only an intermediate step in defining functionality, because it does not tell you whether the mRNA has been translated to a protein or enzyme. When it is possible, direct measurement of metabolites or enzymes produced provides more reliable information on functionality. But when this isn’t possible, or if you need an explorative analysis, RNA analysis can offer important insight.
RNA is very sensitive and easily degraded, so you need to be very careful when sampling, isolating and storing it, to be sure you have the quality and quality to perform the sequencing or qRT-PCR analysis and get accurate information. Also, it’s very timepoint specific: what you will see depends on the moment of sampling. It’s thus more complicated than DNA sampling, and requires specialised skills and knowledge.